Thought Leaders
From Code to Cure: The Next AI Revolution Needs A Hand (And Eyes)

How agentic systems, XR smart glass, and robotics will empower humans—not replace them
We are living through a paradox in artificial intelligence.
On screens, AI is superhuman. Large language models write functional Python code in seconds. Generative systems produce photorealistic images and videos in minutes. Nobel-winning systems like AlphaFold have predicted the structures of nearly all known proteins. The digital victories are stacking up.
Yet in the physical world of biomedical research, the process of discovery remains painstakingly manual. We don’t really feel AI accelerating science or medicine, at least not yet. The numbers reveal the depth of the problem. A landmark Nature survey of over 1,500 scientists found that more than 70% have tried and failed to reproduce another researcher’s experiments. Even more troubling: more than half couldn’t reproduce their own work. In cancer biology specifically, an eight-year reproducibility project found that only 40% of high-impact findings could be replicated and 68% of experiments lacked sufficient documentation to even attempt replication.
This is the dirty secret of modern science: we have a knowledge capture problem, not just a discovery problem. Critical experimental details live in researchers’ heads, not in papers. Protocols drift. Tacit knowledge walks out the door when trainees graduate. AI systems trained on published literature inherit all these gaps.
The fundamental issue is that while an AI can design a novel protein for cancer therapy in a digital simulation, it cannot pick up a pipette to test it. It cannot navigate the messy, unpredictable reality of a wet lab to validate its own hypothesis. It cannot watch an experienced scientist’s hands and learn the subtle techniques that make experiments work.
This “execution gap” is the single largest bottleneck preventing the AI revolution from becoming a medical revolution. While most robotics companies are still busy teaching machines to fold laundry or load dishwashers, they are lagging behind on the truly transformative capabilities of these advancements in fields such as medicine.
To solve this, we must move beyond chatbots and toward AI Co-Scientists, agentic systems that bridge the digital and the physical world, pushing beyond planning and coding and into real-world execution. At Stanford, we are developing LabOS, a digital-physical AI framework that demonstrate how AI agents, Extended Reality (XR) smart glasses, and collaborative robotics can unite to close this loop, transforming scientific experiments into a collaborative conversation between human and machine, while automatically capturing the knowledge that currently gets lost.
The Great Divide: Why AI Needs “Eyes” and “Hands”
Many of the most visible AI wins have happened where the environment is fully digital: code repositories, curated datasets, or simulated benchmarks (where AI competes to run a virtual business or digitally investing in stocks).
Wet labs are different. Biology, and in general, scientific discovery, is a very noisy process. Instruments drift, operators improvise, and “the protocol” often lives partly in people’s heads. The difference between a clean result and a failed run can be a pipetting angle, a vortexing pattern, a reagent substitution, or an incubation that ran 10 minutes long. These contextual details rarely make it into a paper and they are exactly what an AI model needs if it is going to generalize beyond a dataset.
That is why lab-grade AI needs eyes (to perceive what is happening in context), hands (to standardize and safely automate high-variance steps), and memory (to record what actually happened). Without those capabilities, models can recommend what to do, but they cannot reliably translate recommendations into validated physical execution or explain failures when reality diverges from plan.
Beyond Chatbots: From Copilots to Co-Scientists
The term “agentic AI” is sometimes used loosely. In biomedical settings, it should mean something precise: a system that can take a goal (for example, “optimize CRISPR gene editing efficiency while minimizing off-targets”), decompose it into a sequence of tasks, execute those tasks across tools, evaluate outcomes, and adapt the plan under constraints and with auditable decision-making.
This matters because research workflows are not a single model call. They are end-to-end pipelines that span hypothesis formulation, experimental design, data processing, statistical testing, and interpretation. Recent thinking in drug discovery has started to emphasize agentic systems that can scale these pipelines rather than merely accelerate individual steps (for example, Unite.AI’s discussion of agents in small-molecule discovery).
In software engineering, we have already seen early empirical evidence that AI copilots can increase developer throughput. In biomedicine, the analogous opportunity is not just writing code, it is writing and validating protocols, structuring data, monitoring execution, and closing the loop between prediction and measurement, connecting AI to human scientists in labs.
LabOS: When AI Runs on an Operating System for the Labs of Tomorrow
In our work at Stanford on AI4Science, spanning gene-editing copilots such as CRISPR-GPT and AI-XR co-execution systems such as LabOS that help scientists in biomedical and materials science labs, we have been exploring an architectural shift:
1. Designing an end-to-end “lab operating system” that couples a digital (dry) lab to a physical (wet) lab.
The premise is simple: if a lab notebook is the memory of science, then a lab OS should be the execution layer, capturing intent, translating it into actions, observing outcomes, and turning every run into structured knowledge.
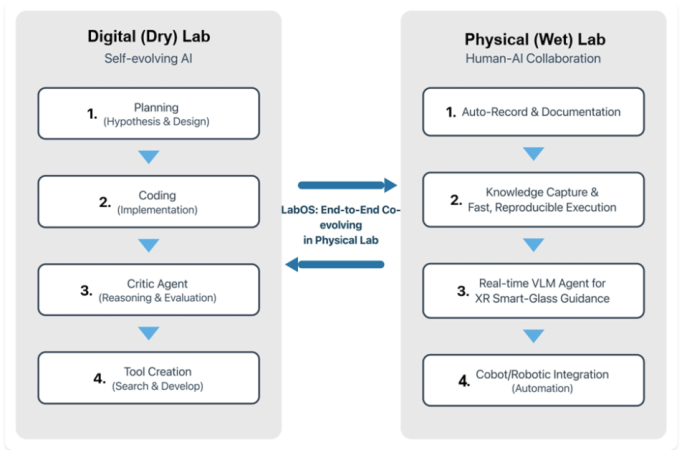
Figure. A conceptual view of LabOS linking a self-evolving digital lab loop (planning, coding, critique, tool creation) with a human-AI wet-lab loop (auto-documentation, knowledge capture, XR guidance, and robotic integration).
2. AI in the Digital Lab – Self-Evolving Planning and Tool-Building
In the digital (dry) lab, we can let AI do what it already does well: search, synthesize, and propose. But we also want it to be self-improving. Not by “hallucinating new science,” but by learning better tools and workflows from feedback.
A practical digital-lab loop can be framed as four recurring stages:
- Planning (hypothesis + design): propose hypotheses, select experimental variables, anticipate confounders, and specify measurable endpoints.
- Coding (implementation): generate or adapt analysis scripts, simulation pipelines, and instrument control templates where appropriate.
- Critic agent (reasoning + evaluation): stress-test assumptions, check statistical power, propose controls, and flag likely failure modes.
- Tool creation (search + develop): when the workflow lacks a component (a parser, a QC routine, a dashboard), build it and add it back to the toolkit.
3. AI in the Physical Lab – AI With “Eyes” (XR glasses) and “Hands” (Robotics)
The physical (wet) lab is where the system either earns trust – or loses it. The goal is not to replace the scientist, but to reduce friction and error while increasing observability.
We view the physical-lab loop as four complementary capabilities:
- Auto-record and documentation: capture actions, timestamps, instrument settings, and deviations automatically, so documentation is not an afterthought.
- Knowledge capture for fast, reproducible execution: convert runs into structured, queryable artifacts (protocol versions, parameter sets, QC outcomes) aligned with data stewardship principles such as FAIR.
- Real-time vision-language guidance via XR smart glasses: use multimodal models to interpret the scene (what the operator is doing, which reagent is in hand) and provide step-by-step guidance and safety checks. AR/XR has already demonstrated value in high-stakes physical workflows like experimental guidance (LabOS, Stanford, Princeton, in collaboration with NVIDIA).
- Cobot/robotic integration for automation: standardize repetitive steps, enable safe hand-offs, and reduce variability. Platforms for simulation-to-real workflows (for example NVIDIA Isaac) and real-time AI processing at the edge such as the smart glass live streaming and human-AI interaction in LabOS are important enabling layers.
This architecture aligns with a broader direction in the field: “self-driving” or autonomous laboratories that combine automation with machine learning to plan the next experiment. What LabOS adds is a tighter human interface layer, so autonomy does not come at the cost of transparency.
Why Lab-Grade AI Isn’t Just “AI on a Dataset”
AI systems for biomedical/scientific super intelligence often look impressive in retrospective evaluation or taking exams, and then underperform in the physical lab. The reason is rarely a single bug. It is usually a mismatch between the model’s assumptions and the lab’s reality.
Three gaps show up repeatedly:
- Context gap: datasets typically omit the contextual variables that operators know matter (temperature excursions, reagent lot numbers, subtle protocol deviations).
- Action gap: many AI systems can recommend what to do, but cannot reliably translate that recommendation into validated physical steps.
- Feedback gap: without structured, high-quality feedback from execution, models cannot learn where they fail – and scientists cannot audit why.
Closing these gaps is less about inventing a new neural network architecture, but more about building the instrumentation, interfaces, and data contracts that make the laboratory legible to machines, and allow AI to see and work with humans.
Trust by Design: Safety and Governance for AI That Can Act
Agentic AI in discovery research does not just raise familiar concerns about accuracy. It introduces new failure modes because it can act. In a lab, action means the potential for waste, harm, or misleading conclusions, especially when experiments feed into clinical hypotheses.
A useful mindset is to treat an AI-enabled lab stack as a socio-technical system that needs assurance. Several existing frameworks help, but they must be translated into lab reality:
- Risk management as a continuous practice: NIST’s AI Risk Management Framework (AI RMF 1.0) provides a practical vocabulary for mapping, measuring, and managing AI risk across the lifecycle.
- Regulatory alignment for medical-adjacent AI: the FDA’s AI/ML Software as a Medical Device (SaMD) work, including its Action Plan and related guidelines, offers a concrete view of what “good practice” looks like when AI impacts patient care.
For gene-editing and other high-consequence domains, governance is already a global conversation. Recommendations are being discussed on human genome editing, to emphasize the need for appropriate oversight mechanisms and responsible governance, such as those put forward by the American Society for Gene and Cell Therapy, or ASGCT. Systems like LabOS should be designed to make compliance easier, not harder.
Checklist: Controls for Safe AI Co-scientists for Scientific Discovery
In our view, a safety-conscious lab OS should implement at least the following designs:
- Provenance by default: every dataset, protocol version, and model output should be traceable to inputs and timestamps.
- Bounded autonomy: the system should have explicit permissions (what it can do without confirmation) and escalation rules (when it must ask).
- Human override and graceful degradation: when sensors or data streams fail, or uncertainty is high, the system should fall back to a safer, simpler mode.
- Continuous validation: in-silico predictions should be paired with physical-lab validation; physical lab runs should include QC gates before conclusions propagate downstream back to the AI models/agents in the digital world.
- Security and dual-use awareness: protect lab infrastructure from tampering.
Empower Humans Everywhere: Can AI Co-Scientists Level the Playing Field?
One of the most compelling promises of an AI-XR “co-scientist” is not just speed for elite institutions, it is accessibility for everyone. Consider what currently limits smaller labs, startups, and remote/rural/regional clinics:
- Limited access to specialized expertise for gold-standard protocols and instruments.
- Higher relative cost of training, mistakes, and rework.
- Fragmented tooling: notebooks, spreadsheets, instrument logs, and analysis scripts rarely connect cleanly.
A system that can guide execution in context (through XR glass), capture what happened automatically, and suggest the next best step based on prior runs could make advanced assays more repeatable across sites. In principle, it could also support distributed clinical research where protocols must be executed consistently, even when resources vary.
Timeline: When Does Every Scientist and Clinician Get a Co-Scientist?
In short, we are closer than many think for some high-value, high-frequency tasks (such as producing a drug reliably in the lab) and farther than most demos imply for others (such as AI completely solve big problems like cancer or Alzheimer’s). A realistic roadmap looks like this:
- Near term (within 1 year): Workflow copilots that reduce administrative load: protocol drafting, literature synthesis, analysis templates, and automated QC reports. The limiting factor is integration, not model capability.
- Mid term (1-2 years): Co-execution systems in the lab: XR glass guidance, automated documentation, and selective robotics for high-variance steps. Trust will depend on audit trails and tight human-in-the-loop design.
- Longer term (3+ years): Cross-domain co-researchers that connect discovery to translation: linking lab data to clinical endpoints, monitoring safety signals, and helping design trials – while complying with regulatory and ethical expectations.
From Code to Cure: The Path Forward to 1000x Scientific Discovery
LabOS is one attempt to answer a simple question: what if an experiment could be run as a conversation, where intent, execution, and evidence are connected end-to-end? If we build these systems well, they can help address the translation gap that slows biomedicine and many physical science disciplines (e.g. material science). If we build them poorly, they will amplify irreproducibility and create new safety risks.
The most important work in the next few years will be foundational: standardized data and device interfaces via building the operating system (much like iOS runs all type of apps), building AI benchmarks that include execution and uncertainty (like the LabSuperVision benchmark in LabOS), and starting on deployment in real world that encourage innovation while protecting patients and research integrity.
For researchers and clinicians, the question is not whether AI will enter the lab. It already has. The question is whether we will integrate it as a collection of disconnected tools or as a trustworthy, auditable system designed for the realities of biomedical science.
Suggested reading and sources
- Reproducibility survey (Nature, 2016): https://www.nature.com/articles/533452a
- CRISPR-GPT peer-reviewed article (Nature Biomedical Engineering): https://www.nature.com/articles/s41551-025-01463-z
- Stanford Medicine news on CRISPR-GPT (2025): https://med.stanford.edu/news/all-news/2025/09/ai-crispr-gene-therapy.html
- LabOS preprint (arXiv): https://arxiv.org/abs/2510.14861
- LabOS and LabSuperVision benchmark website: https://ai4lab.stanford.edu
- NIST AI Risk Management Framework (AI RMF 1.0): https://nvlpubs.nist.gov/nistpubs/ai/NIST.AI.100-1.pdf
- FDA overview of AI/ML Software as a Medical Device: https://www.fda.gov/medical-devices/software-medical-device-samd/artificial-intelligence-software-medical-device
- FAIR data principles (Scientific Data, 2016): https://doi.org/10.1038/sdata.2016.18
- Unite.AI on bottlenecks in small-molecule drug discovery: https://www.unite.ai/how-ai-is-breaking-down-the-bottlenecks-in-small-molecule-drug-discovery/
- Unite.AI on AI and robotic surgery: https://www.unite.ai/how-ai-is-ushering-in-a-new-era-of-robotic-surgery/












