Ângulo de Anderson
Cada Vez Mais, o HIPAA Não Consegue Impedir que a IA Desanonymize Dados de Pacientes

Even after hospitals strip out names and zip codes, modern AI can sometime still work out who patients are. Great news for insurance companies; not so much for healthcare recipients.
New research from New York University finds that US patients’ medical notes, stripped of names and other HIPAA identifiers, can expose patients to re-identification. By training AI language models on a large corpus of real world, uncensored patient records, identity-defining details remain – in some cases allowing a patient’s neighborhood to be inferred from diagnosis alone.
The new study places this risk in the context of a lucrative market in de-identified health data, where hospitals and data brokers routinely sell or license scrubbed clinical notes to pharmaceutical firms, insurers, and AI developers.
The authors of the new study challenge even the very concept of ‘de-identification’, enshrined in the patient protections established by HIPAA after Massachusetts Governor William Weld has his medical data de-anonymized in 1997:
‘[Even] under perfect Safe Harbor compliance, “de-identified” notes remain statistically tethered to identity through the very correlations that confirm their clinical utility. The conflict is structural instead of technical.’
The researchers contend that current, HIPAA-compliant de-identification frameworks leave two backdoors available for ‘linkage attacks’:
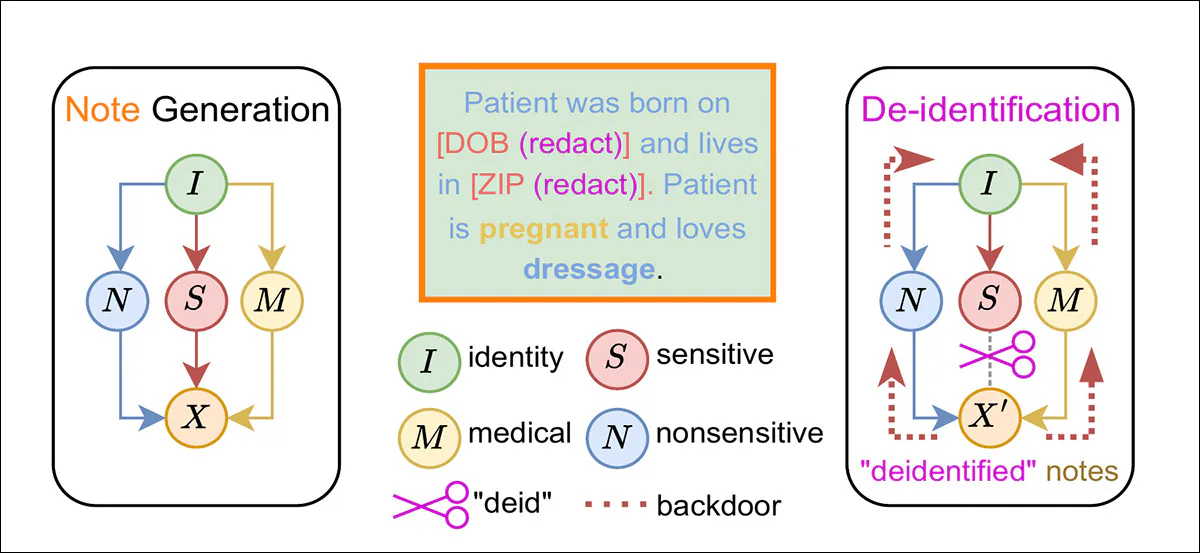
From the new paper, a causal diagram illustrating how HIPAA-style de-identification removes explicit sensitive attributes while leaving identity-linked correlations intact, allowing patient identity to be inferred through non-sensitive and medical information. Source
In the example above we see not only that the patient is pregnant – the lowest-hanging fruit in de-identification, since it establishes biological gender unequivocally – but also that she likes a pastime not associated with lower-income groups, according to the researchers:
‘Although the protected attributes (DOB and ZIP code) are redacted, we can still infer that the patient is an adult female based on the pregnancy, and resides in an affluent neighborhood given the hobby of dressage.’
In one experiment, even after patient identifiers were stripped, more than 220,000 clinical notes from 170,000 NYU Langone patients still carried enough signal to allow demographic traits to be inferred.
Drilling Down
A BERT-based model was fine-tuned to predict six attributes from the de-identified records, and, the paper notes, exceeded random-chance guesses with as few as 1,000 training examples. Biological sex was recovered with over 99.7% accuracy, and even weaker cues such as the month the notes were taken were predicted at above-chance levels.
For experimental purposes, those inferred traits were then used in a linkage attack against the Langone database, producing a maximum unique re-identification risk of 0.34% – roughly 37 times higher than a simple majority-class baseline. Applied to the US population, this attack alone would de-identify 800,000 patients.
The authors frame the problem as a ‘paradox’, because what is left behind in HIPAA-compliant de-identified patient records is clearly a viable basis for de-identification attacks:
‘[The] vast majority of re-identification risk stems not from Protected Health Information, but from the non-sensitive and medical content we deem safe to share.’
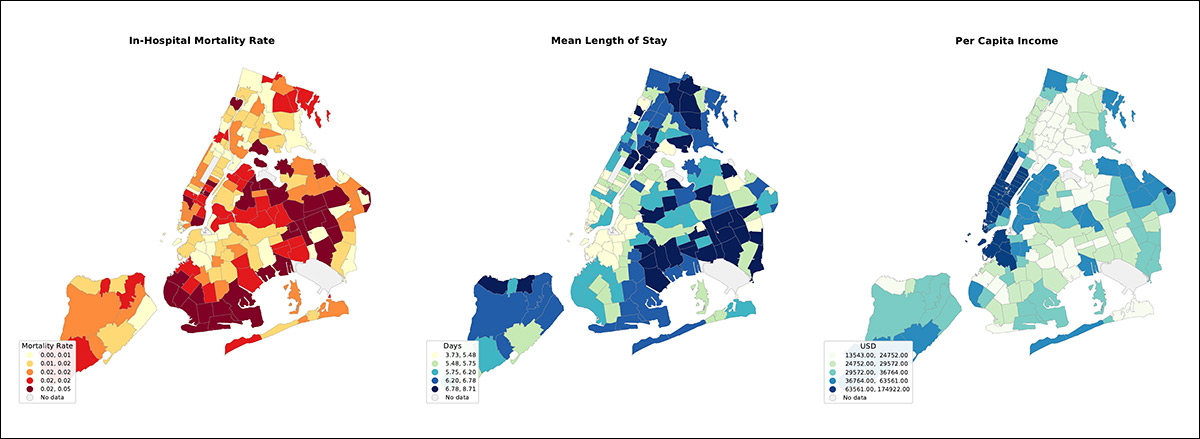
Maps of New York City boroughs showing differences in hospital death rates, average length of stay, and income levels, illustrating how health outcomes and wealth vary by area and can leave location-linked clues even in de-identified medical notes. Please refer to source paper for additional examples
The paper argues that HIPAA’s Safe Harbor rules no longer work the way policymakers intended: removing 18 identifiers may satisfy the letter of the law, but according to the authors, it does not prevent identity from being inferred by current language models. They frame the system itself as built on outdated assumptions regarding what LLMs can and cannot infer from ordinary medical text.
The work also suggests that those likely to benefit from the weaknesses stated are large corporations related to medical insurance, rather than conventionally-defined criminal entities (such as hackers, blackmailers, or social engineers)*:
‘The persistence of Safe Harbor despite known limitations is not an oversight but a feature of a system optimized for data liquidity rather than patient protection. De-identified clinical notes represent a multi-billion dollar market, creating structural disincentives for healthcare institutions to adopt privacy-preserving alternatives that might reduce data utility or require costly infrastructure investments.
‘There is an urgency to carefully investigate, understand and address this disincentive.’
This is a position paper, with no clear answers offered; however, the authors suggest that research into de-identification should pivot towards social contracts and legal consequences of breach, rather than technical solutions (arguably the same approach used by the DMCA to restrict copying of IP-protected work, when technical solutions failed).
The new paper is titled Paradox of De-identification: A Critique of HIPAA Safe Harbour in the Age of LLMs, and comes from four researchers at New York University, in association with NYU Langone hospital.
Method
To test their theory, the authors developed a two-stage linkage attack using 222,949 identified clinical notes from 170,283 patients treated at NYU Langone, with all notes partitioned by patient into 80% training, 10% validation, and 10% test splits, to prevent cross-contamination.
For context, this collection is 3.34 times larger than the MIMIC-IV dataset, the largest publicly-available Electronic Health Records (EHR) collection. For privacy reasons, the Langone dataset will not be made available in any form, though users can experiment with the project’s principles via a GitHub repo that generates synthetic data.
Six demographic attributes were curated to approximate the classic re-identification trio identified in an influential prior work: biological sex; neighborhood; note year; note month; area income; and insurance type:
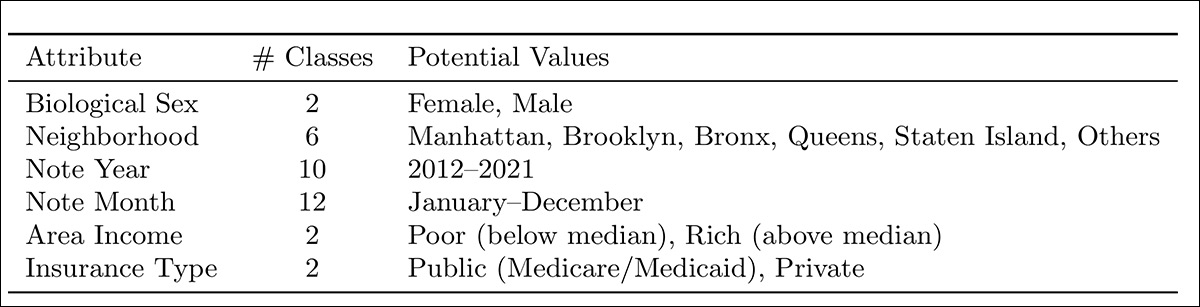
Demographic attributes inferred from de-identified NYU Langone clinical notes, comprising biological sex, neighborhood, note year, note month, area income, and insurance type, selected to approximate the unique identifier trio described in ‘Simple Demographics Often Identify People Uniquely’.
The notes were de-identified using UCSF philter before modeling.
A BERT-base-uncased model with 110 million parameters, pre-trained on general-domain text to avoid prior exposure to clinical data, was fine-tuned separately for each attribute, using eight NVIDIA A100 GPUs with 40GB memory, or H100 GPUs, with 80GB memory, for up to ten epochs. Optimization used AdamW, with a learning rate of 2×10−5, and an effective batch size of 256
Generalization on the held-out test set was evaluated using Accuracy and weighted ROC-AUC, the latter chosen to account for class imbalance across attributes.
To make the attack more realistic, the model’s predictions were not treated as single definitive answers. Instead, for each attribute, the top k most likely values were kept, and the patient database filtered to include anyone who matched those predicted traits. This produced a shortlist of possible identities for each note, rather than a single guess.
Risk Assessment
Re-identification risk was then calculated in two stages: measuring how often the real patient appeared inside that shortlisted group; and estimating the chance of selecting the correct person from within that group.
Because the last step assumed someone simply picked a name at random from the possible matches, the reported number is a cautious estimate, and a determined attacker could likely do better.
The experiment assumed access to the full patient population in the external database. This reflects a worst-case but realistic scenario in which a large institution or data broker, with broad coverage of patient records, attempts the linkage, rather than an individual acting with limited information, further reinforcing the nature of the threat that the authors are addressing in the work.












